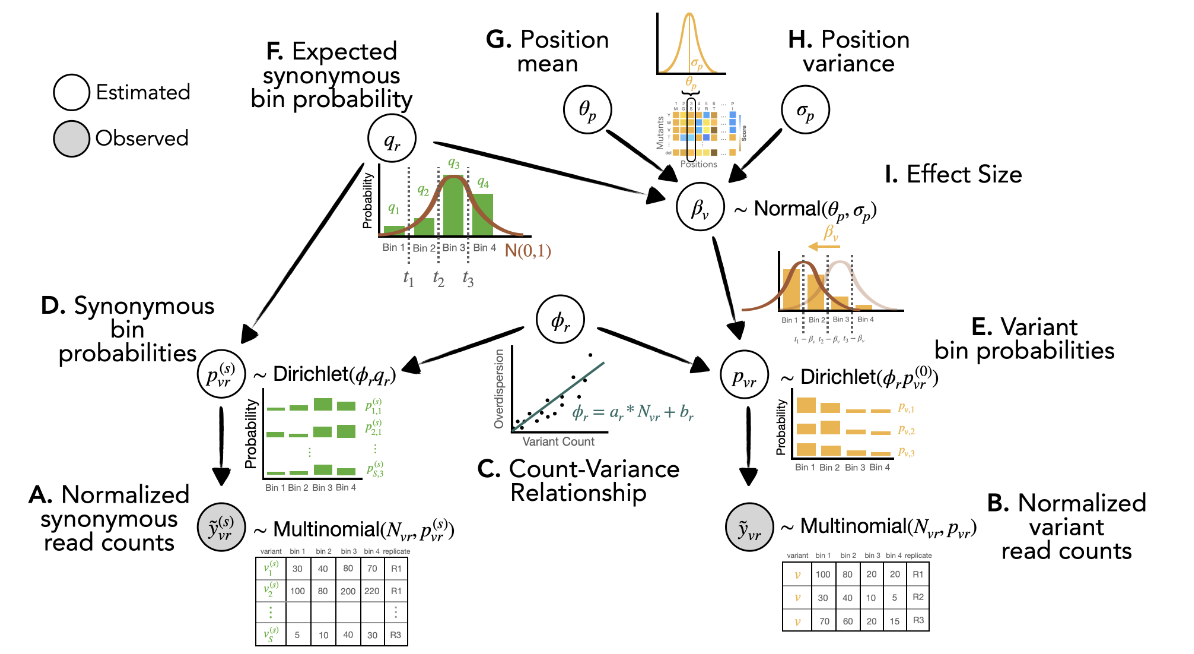
Cosmos: A Position-Resolution Causal Model for Direct and Indirect Effects in Protein Functions
Rao J, Wang M, Howard MK, Coyote-Maestas W*, Pimentel H*
Biorxiv,
2025
Accurate variant effect estimation in FACS-based deep mutational scanning data with Lilaces
Freudenberg J, Rao J, Howard MK, Macdonald CB, Greenwald NF, Coyote-Maestas W*, Pimentel H*
Biorxiv,
2025
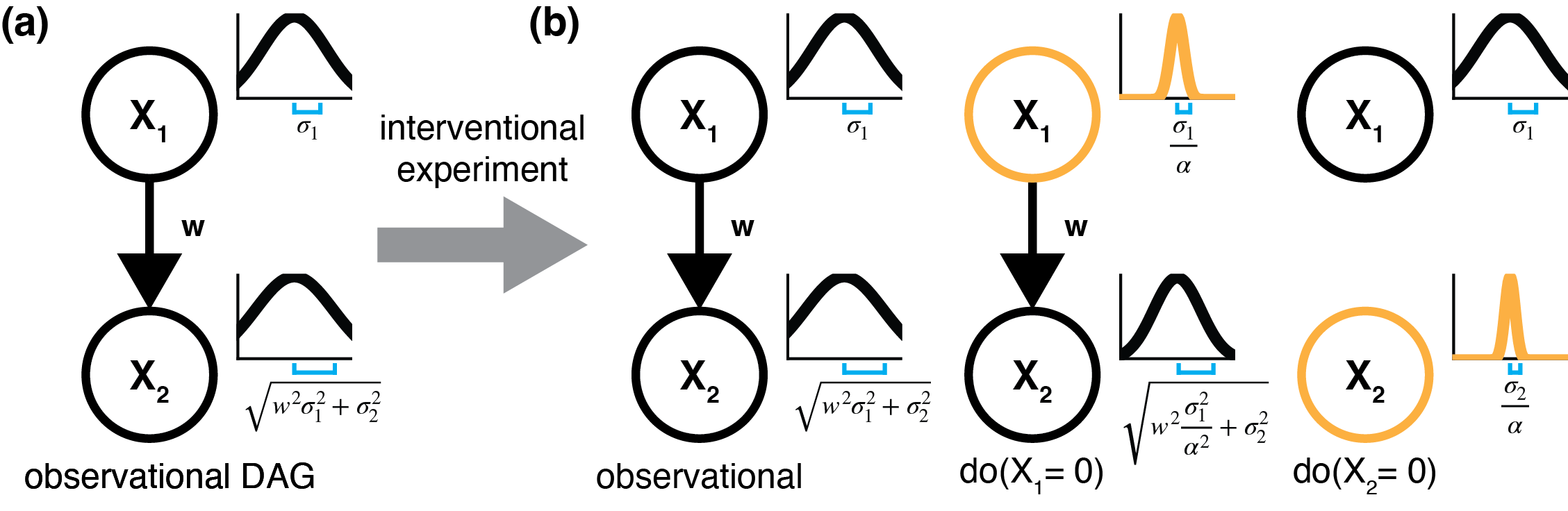
dotears: Scalable, consistent DAG estimation using observational and interventional data
Xue A, Rao J, Sankararaman S*, Pimentel H*
iScience,
2025
Rosace-AA: Enhancing Interpretation of Deep Mutational Scanning Data with Amino Acid Substitution and Position-Specific Insights
Rao J*, Wang M, Howard MK, Macdonald CB, Fraser JS, Coyote-Maestas W*, Pimentel H*
Biorxiv,
2025
Rosace: a robust deep mutational scanning analysis framework employing position and mean-variance shrinkage
Rao J, Xin R, Macdonald CB, Howard M, Estevam GO, Yee SW, Wang M, Fraser JS, Coyote-Maestas W*, Pimentel H*
Genome Biology,
2024